xbreed: An R package for genomic simulation of purebred and crossbred populations
|
|
|
- Jewel Holland
- 6 years ago
- Views:
Transcription
1 xbreed: An R package for genomic simulation of purebred and crossbred populations Version Hadi Esfandyari, Anders Christian Sørensen 03 April 2017 Contents Introduction Workflow Validation of simulated genomes Heterozygosity and Linkage disequilibrium Allele frequency distribution Evaluation of simulated genomes Implementation and efficiency Examples Example 1: Make historical population Example 2: Simulation of an additive trait Example 3: Simulation of a trait with additive and dominance effects Example 4: Multiple recent populations and LD Example 5: Two-way crossbreeding Example 6: Three-way crossbreeding Example 7: Four-way crossbreeding Acknowledgments References
2 Introduction Simulation has and will continue to play an important role in the study of genomic selection. xbreed is a simulation tool for purebred and crossbred genomic data as well as pedigree and phenotypes. It can be used for simulation of a population with flexible genome structures and trait genetic architectures. xbreed can also be used to evaluate breeding schemes and generate genetic data to test statistical tools. Furthermore, the package is handy, intuitive and has good performance properties in execution time and memory usage, even when handling large amounts of genetic data. This vignette intends to explain in detail how xbreed package works for the desired applications, and it includes illustrated explanations and easily reproducible examples. In conclusion, xbreed would be a useful tool for the methodological and theoretical studies in the population and quantitative genetics and breeding. Workflow Simulation in xbreed is basically carried out in two steps. In the first step, historical generations are simulated to create desirable level of LD and in the second step, recent population structures are generated, which can be as simple as a purebred line or a four-way crossbreed population. xbreed allows for a wide range of parameters to be incorporated in the simulation models in order to produce appropriate simulation data. make_hp is the central function in the package (Figure 1). This function is used to create historical population and genomic structure of the trait of interest. A wide range of parameters can be specified for simulating the genome, such as: number of chromosomes, markers and QTL, location of markers and QTL, mutation rate and initial allelic frequencies. This flexibility permits for a wide variety of genetic architectures to be considered. As an example, user can simulate SNP chip genotypes data mimicking the real genotypic data of different livestock species. Other functions in the package are dependent to the function make_hp. Function sample_hp is used to sample individuals from the last generation of historical population. This function can be used multiple times to sample individuals from the last generation of historical population to create recent populations. Function make_hp works on the output of the function sample_hp and also itself (Figure 1). This means that, once a recent population (e.g. RP1) is created by sample_hp, function make_rp can be used, to create the second recent population (RP2) based on the data of RP1. Features of the functions sample_hp and make_rp are the same as both can create recent populations. However, the difference between these two functions is that, for function sample_hp, male and female founders are always from the last generation of historical population, whereas, in function make_rp, male and female founders can be selected from any generation of the population created by sample_hp or itself. In othe words, function make_hp can get the output of itself as input. See examples for more clarification. Function xbreed is used for crossing between populations. These populations can be the ones created by functions make_rp and sample_hp (Figure 1). Also, if user would like to have 2
3 multi-way crossbreeding schemes such as three-way or four-way crossbreeding, then function xbreed can be used to get the output of itself as input data in order to create the multi-way crossbreed populations. Simulations of two-way and multi-way crossbreeding schemes are presented in the example section. Figure 1: Diagram of simulation workflow. make_hp: Function to make historical population, sample_hp: Function to make a population by sampling from historical population, make_rp: Function to make a recent population, xbreed: Function to make a crossbred population. Validation of simulated genomes Heterozygosity and Linkage disequilibrium A number of arbitrary assumptions are made during the simulation of genomes, which makes it necessary to confirm that characteristics of the simulated data are similar to expectations. Equations for LD [r 2 (Hill and Robertson 1968)] and heterozygosity given some population parameters have been described in the literature. Deterministic predictions for these parameters may not exist for complex population histories, which may involve expansion or reductions in N e. However, in such cases simulation programs can still be evaluated using a simple population history before moving on to more complex models. The expected heterozygosity of loci, H e, for a given effective population size, N e, and mutation rate, u, is E(H e ) 4N e u/[4n e u + 1] (Kimura and Crow 1964). Under infinite-allele model, mutation creates new unique allele that never existed in the population. For finite-allele model, mutation does not create novel allele but rather modifies an allele to another allele within the allelic space with the same 3
4 probability. Recurrent mutation can be reversible. At equilibrium H e is approximately, E(H e ) 1 where k is the number of possible alleles. [ ] 1 + (4Ne u/k 1) 1 + (4N e uk/k 1) Similarly, expected values for LD have been described for scenarios without mutation, E(r 2 ) 1/[1 + 4N e c] (Sved 1971), and with mutation, E(r 2 ) 1/[2 + 4N e c] (Tenesa et al. 2007), where c is the recombination rate. Hudson (1985) has shown that expectations are met only when loci with allele frequency, 0.05 are removed from LD calculations. Allele frequency distribution Considering a finite population and the recurrent mutation of two alleles, Wright (1937) shows that the probability of the allele frequency (p) at mutation-drift equilibrium can be obtained by the asymptotic formula: prob(p) p 4Neu 1 (1 p) 4Nev 1 where u and v are the mutation rates of two alleles. This formula indicates the allele frequency distribution at mutation-drift equilibrium depends only on N e, u and v. Under the assumption that u = v, when 4N e u is less than 1, close to 0, and greater than 1, the allele frequencies are expected to exhibit a U-shaped, uniform and convex distribution, respectively. Evaluation of simulated genomes In the following we assessed the characteristics of the genome simulated by xbreed. An historical population was simulated for 2000 generations of random mattings with an effective population size (N e ) of 100 (50 males and 50 females). The simulated genome consisted of one chromosome with a length (L) of 1 Morgan, containing 2000 randomly spaced SNP loci. The initial minor allele frequency (laf) of all SNPs was assumed to be 0, meaning all individuals were completely homozygous for the same allele in the first generation of the historical population. Mutation occurred at a rate (mutr) of per locus meiosis and involved the switching from one allele to another. Recombinations were sampled from a Poisson distribution with a mean of 1 per Morgan and were then randomly placed along the chromosome. The code used for assessing quality of simulated genome is in Example the mean heterozygosity and r 2 of all replicates was compared to expected values (Table 1). Table 1. Actual and expected Heterzygosity, H e, and LD between adjacent loci, r 2, in simulated data. Expected Observed He LD(r2)
5 Decay of average r 2 over distance in the last generation of historical population is in Figure 2. To determine the decay of LD with increasing distance between SNPs, the average r 2 was expressed as a function of distance between SNPs. SNP pairs were grouped by their pairwise distance into intervals of 1 cm, starting from 0 up to 100 cm. The average r 2 for SNP pairs in each interval was estimated as the mean of all r 2 within that interval. Figure 2: Decay of average r2 over distance The allele frequency distributions in the last generation of the historical population are in Figure 3. Based on the simulation parameters defined above, the value for 4N e u was 0.1 that is smaller than one. At mutation-drift equilibrium, for values of 4N e u smaller than 1, a U-shaped distribution is expected. Also, we considered a case that 4N e u was greater than 1. To do so, we increased mutation rate (mutr) to When 4N e u is larger than 1, normal distribution of allele frequencies is expected, respectively (Wright, 1937). Implementation and efficiency The program is written in R, however time-consuming parts of each function in the package are in Fortran. So, compared to routine packages in R, xbreed is efficient in terms of computing time. The computing time and RAM demanding on a mediocre PC, with 2.3 GHz CPU, 4 GB RAM is 1.51 minutes and 5 Mb, respectively, for simulating 10K SNP panel, 5
6 Figure 3: Distribution of allele frequencies in the last generation of the historical population for 4Neu<1 (a) and 4Neu>1 (b) historical generations with N e = 100. The most important parameters affecting CPU time and memory are population size and the number of loci (markers and QTL). However, depending on the simulation scenario, other parameters such as gebv(c) as selection criteria which demands training on each generation of simulation, and saving huge outputs, etc. might become a computational bottleneck. Nonetheless, xbreed provides the user with enough options to manage the outputs or turn off unwanted computations. For example, one may optionally save genotypes only for last few generations avoiding large outputs. Examples Examples below demonstrate some main features of the package. The mechanism to create markers, population structure, and trait phenotypes are detailedly proposed. The idea was to show some basic breeding schemes, however more complex simulation structures are possible according to the user desire. Example 1: Make historical population Assessing quality of simulated genome for a population of N e =100 for 1000 historical generations. library(xbreed) #Genome parameters genome<-data.frame(matrix(na, nrow=1, ncol=6)) 6
7 names(genome)<-c("chr","len","nmrk","mpos","nqtl","qpos") genome$chr[1]<-c(1) #Chromosome id genome$len[1]<-c(100) #Chromosome length in cm genome$nmrk[1]<-c(2000) #Number of markers genome$mpos[1]<-c('rnd') #Position of markers genome$nqtl[1]<-c(2) #Number of qtl genome$qpos[1]<-c('rnd') #Position of qtl genome hp<-make_hp(hpsize=100,ng=1000,h2=0.3,d2=0,phen_var=1,genome=genome,mutr=2.5*10**-4,laf=1) # Expected Heterozygosity according to (Kimura and Crow 1964) mutr<-2.5*10**-4 ne<-100 k<-2 Fneu<-4*ne*mutr Expected_het1<-1-((1+((Fneu)/(k-1)))/(1+((Fneu*k)/(k-1)))) Expected_het2<-Fneu/(1+Fneu) Expected_het1 Expected_het2 # Observed Heterozygosity het_observed<-mean(2*(hp$freqmrk[,3]*hp$freqmrk[,4])) het_observed # Mean r2 and LD decay mat<-hp$hp_mrk[,-1] LD<-calc_LD(mat=mat,MAF=0.1,method='adjacent',LD_summary=TRUE) LD$Mean_r2 linkage_map<-hp$linkage_map_mrk[,3] LD<-calc_LD(mat=mat,MAF=0.10, method='pairwise',ld_summary=true, linkage_map=linkage_map,interval=0.5) LD$ld_decay Example 2: Simulation of an additive trait Simulation of a recent population following historical population. Simulation of an additive trait with h 2 =0.3. A genome consisting of three chromosomes with different parameters. 7
8 # Genome consisted of 3 chromosomes genome<-data.frame(matrix(na, nrow=3, ncol=6)) names(genome)<-c("chr","len","nmrk","mpos","nqtl","qpos") genome$chr<-c(1:3) genome$len<-c(146,126,116) genome$nmrk<-c(1000,2000,500) genome$mpos<-c('even','rnd','rnd') genome$nqtl<-c(80,100,90) genome$qpos<-c('rnd','even','rnd') genome # CREATE HISTORICAL POPULATION hp<-make_hp(hpsize=200,ng=500,h2=0.3,phen_var=1,genome=genome,mutr=5*10**-3,sel_seq_qtl=0.1,sel_seq_mrk=0.05,laf=0.5) # # MAKE RECENT POPULATION # Select 50 males randomly as founders of recent population Male_founders<-data.frame(number=50,select='rnd') # Select 100 females as founders of recent population based on phenotype Female_founders<-data.frame(number=100,select='phen',value='h') # Selection of animals in each generation of recent population # Selection of 60 sires and 200 dam # Selection criteria is "tbv" for sires and "gebv" for dams Selection<-data.frame(matrix(NA, nrow=2, ncol=3)) names(selection)<-c('number','type','value') Selection$Number[1:2]<-c(60,200) Selection$type[1:2]<-c('tbv','gebv') Selection$Value[1:2]<-c('h','h') Selection # Training parameters for the estimation of marker effects. Training<-data.frame(matrix(NA, nrow=1, ncol=8)) names(training)<-c('size','sel','method','niter','burnin','thin','save','show') Training$size<-200 Training$sel<-'min_rel_mrk' Training$method<-'BRR' Training$nIter<-1000 Training$burnIn<-100 Training$thin<-5 Training$save<-'bayes' Training$show<-TRUE Training # Save output of simulation for "data", "qtl" and "freq_mrk" 8
9 sh_output<-data.frame(matrix(na, nrow=3, ncol=3)) names(sh_output)<-c("data","qtl","freq_mrk") sh_output[,1]<-c(0,3,4) # Save data for generations 0,3,4 sh_output[,2]<-c(1,2,4) # Save qtl genotype for generations 1,2,4 sh_output[,3]<-c(3,4,5) # Save marker frequencies for generations 3,4,5 sh_output # CREATE RECENT POPULATION my_hp<-sample_hp(hp_out=hp,male_founders= Male_founders,Female_founders=Female_founders, ng=4,selection=selection,training=training, litter_size=5,saveat="my_pop",sh_output=sh_output,display=true) # SOME OUTPUT my_hp$summary_data my_hp$output[[1]]$data # Data for base generation (0) my_hp$output[[2]]$freqmrk # Marker frequncy for generation 1 head(my_hp$output[[4]]$qtl) # QTL genotye of individuals for generation 3 my_hp$trait # Tait specifications Example 3: Simulation of a trait with additive and dominance effects Simulation of a trait with both additive and dominance effects with h 2 =0.3 and dominance variance of 0.1. Two recent populations are created. In the first recent population (RP1) founders are from the last generation of historical population. In the second recent population (RP2), male and female founders are from different generations of RP1. Simulated genome is 10k SNP panel with 20 qtl on each chromosome. # CREATE HISTORICAL POPULATION # 10k SNP panel genome<-data.frame(matrix(na, nrow=30, ncol=6)) names(genome)<-c("chr","len","nmrk","mpos","nqtl","qpos") genome$chr<-c(1:30) genome$len<-rep(100,30) genome$nmrk<-rep(333,30) genome$mpos<-rep('rnd',30) genome$nqtl<-rep(20,30) genome$qpos<-rep('even',30) genome hp<-make_hp(hpsize=200,ng=500,h2=0.3,d2=0.1,phen_var=1,genome=genome,mutr=5*10**-4,sel_seq_qtl=0.05,sel_seq_mrk=0.05,laf=0.5) 9
10 # # MAKE RECENT POPULATION ONE (RP1) # Select 50 males 100 females randomly as founders of recent population Male_founders<-data.frame(number=50,select='rnd') Female_founders<-data.frame(number=100,select='rnd') # Selection of animals in each generation of recent population # Selection of 70 sires and 110 dam # Selection criteria is "rnd" for both sires and dams Selection<-data.frame(matrix(NA, nrow=2, ncol=2)) names(selection)<-c('number','type') Selection$Number[1:2]<-c(70,110) Selection$type[1:2]<-c('rnd','rnd') Selection # Save "data" and "marker" for first and last generation of RP1 sh_output<-data.frame(matrix(na, nrow=2, ncol=2)) names(sh_output)<-c("data","marker") sh_output[,1]<-c(1,5) # Save data for generations 1 and 5 sh_output[,2]<-c(1,5) # Save marker genotype for generations 1 and 5 sh_output RP_1<-sample_hp(hp_out=hp,Male_founders= Male_founders,Female_founders=Female_founders, ng=5,selection=selection,litter_size=5,saveat="rp1", sh_output=sh_output,display=true) # # MAKE RECENT POPULATION TWO(RP2) # Select founders for RP2 # Select 45 males based on 'tbv' from generation 4 of RP1. Males<-data.frame(number=45,generation=4,select='tbv',value='h') # Select 80 females based on 'phen' from generation 5 of RP1. Females<-data.frame(number=80,generation=3,select='phen',value='l') # Selection of animals in each generation of RP2 # Selection of 40 sires and 80 dam # Selection criteria is "tbv" for sires and "gebv" for dams Selection<-data.frame(matrix(NA, nrow=2, ncol=3)) names(selection)<-c('number','type','value') Selection$Number[1:2]<-c(40,80) Selection$type[1:2]<-c('tbv','gebv') Selection$Value[1:2]<-c('h','h') Selection # Training parameters Training<-data.frame(matrix(NA, nrow=1, ncol=3)) names(training)<-c('size','sel','method') Training$size<
11 Training$sel<-'rnd' Training$method<-'BL' Training # Save all data for the last two generations of RP2 rp2_output<-data.frame(matrix(na, nrow=2, ncol=6)) names(rp2_output)<-c('data','qtl','marker','seq','freq_qtl','freq_mrk') rp2_output[,1]<-c(3,4) rp2_output[,2]<-c(3,4) rp2_output[,3]<-c(3,4) rp2_output[,4]<-c(3,4) rp2_output[,5]<-c(3,4) rp2_output[,6]<-c(3,4) rp2_output RP_2<-make_rp(sh_out=RP_1,Male_founders=Males, Female_founders=Females,Selection=Selection, ng=4,litter_size=4,saveat='rp2',training=training, rp_output=rp2_output) # Some Output RP_2$summary_data RP_2$output[[4]]$data # Data for base generation 3 RP_2$output[[3]]$freqQTL # QTL frequncy for generation 2 head(rp_2$output[[4]]$mrk) # Marker genotye of individuals for generation 3 Example 4: Multiple recent populations and LD Create multiple recent populations by function make_rp and measurement of LD for each chromosome in the last recent population. # CREATE HISTORICAL POPULATION # Genome consisted of 3 chromosomes genome<-data.frame(matrix(na, nrow=3, ncol=6)) names(genome)<-c("chr","len","nmrk","mpos","nqtl","qpos") genome$chr<-c(1:3) genome$len<-c(110,123,109) genome$nmrk<-c(980,1000,1500) genome$mpos<-c('rnd','rnd','rnd') genome$nqtl<-c(80,100,90) genome$qpos<-c('rnd','rnd','rnd') genome hp<-make_hp(hpsize=100,ng=300,h2=0.3,d2=0.1,phen_var=1 11
12 ,genome=genome,mutr=5*10**-4,sel_seq_qtl=0.05,sel_seq_mrk=0.05,laf=0.5) # # MAKE RECENT POPULATION ONE (RP_1) BY FUNCTION sample_hp # Selection of founders for RP_1 Male_founders<-data.frame(number=50,select='rnd') Female_founders<-data.frame(number=50,select='rnd') # Selection scheme in RP_1 Selection<-data.frame(matrix(NA, nrow=2, ncol=2)) names(selection)<-c('number','type') Selection$Number[1:2]<-c(30,60) Selection$type[1:2]<-c('rnd','rnd') Selection RP_1<-sample_hp(hp_out=hp,Male_founders= Male_founders,Female_founders=Female_founders, ng=5,selection=selection,litter_size=4) # # MAKE RECENT POPULATION TWO (RP_2) BY FUNCTION make_rp # Selection of founders from RP_1 Males<-data.frame(number=45,generation=5,select='tbv',value='h') Females<-data.frame(number=80,generation=5,select='phen',value='l') # Selection scheme in RP_2 Selection<-data.frame(matrix(NA, nrow=2, ncol=3)) names(selection)<-c('number','type','value') Selection$Number[1:2]<-c(40,80) Selection$type[1:2]<-c('phen','phen') Selection$Value[1:2]<-c('h','h') Selection RP_2<-make_rp(sh_out=RP_1,Male_founders=Males, Female_founders=Females,Selection=Selection, ng=7,litter_size=4) # # MAKE RECENT POPULATION THREE (RP_3) BY FUNCTION make_rp # Selection of founders from RP_2 Males<-data.frame(number=40,generation=3,select='phen',value='h') Females<-data.frame(number=70,generation=7,select='phen',value='h') # Selection scheme in RP_3 Selection<-data.frame(matrix(NA, nrow=2, ncol=3)) names(selection)<-c('number','type','value') Selection$Number[1:2]<-c(40,80) Selection$type[1:2]<-c('tbv','phen') Selection$Value[1:2]<-c('h','h') Selection 12
13 RP_3<-make_rp(sh_out=RP_2,Male_founders=Males, Female_founders=Females,Selection=Selection, ng=4,litter_size=5) # # Measurment of LD for the last generation # of RP_3 for each chromosomes # Extracting No of markers in each chromosome linkage_map<-rp_3$linkage_map_mrk No_mrk_in_each_chr<-c() for (i in 1:3){ a<-subset(linkage_map,linkage_map[,2]==i) No_mrk_in_each_chr[i]<-length(a[,3]) } No_mrk_in_each_chr # Measurments of LD in Chromosome 1 chr<-1 generation<-4 # Generation 4 linkage_map<-rp_3$linkage_map_mrk linkage_map<-subset(linkage_map,linkage_map[,2]==chr) linkage_map<-linkage_map[,3] x<-no_mrk_in_each_chr[chr] mat<-rp_3$output[[generation+1]]$mrk mat<-mat[,-c(1,2)] mat<-mat[,1:(x*2)] # Mean r2 rld<-calc_ld(mat=mat,maf=0.1,method='adjacent') # LD decay rld<-calc_ld(mat=mat,maf=0.1,method='pairwise', LD_summary=TRUE,linkage_map=linkage_map,interval=1) rld$ld_decay # Measurments of LD in Chromosome 2 chr<-2 generation<-4 # Generation 4 linkage_map<-rp_3$linkage_map_mrk linkage_map<-subset(linkage_map,linkage_map[,2]==chr) linkage_map<-linkage_map[,3] C1<-No_mrk_in_each_chr[1]*2 C2<-(No_mrk_in_each_chr[1]+No_mrk_in_each_chr[2])*2 mat<-rp_3$output[[generation+1]]$mrk 13
14 mat<-mat[,-c(1,2)] mat<-mat[,(c1+1):c2] # Mean r2 rld<-calc_ld(mat=mat,maf=0.1,method='adjacent') # LD decay rld<-calc_ld(mat=mat,maf=0.1,method='pairwise', LD_summary=TRUE,linkage_map=linkage_map,interval=1) rld$ld_decay # Measurments of LD in Chromosome 3 chr<-3 generation<-4 # Generation 4 linkage_map<-rp_3$linkage_map_mrk linkage_map<-subset(linkage_map,linkage_map[,2]==chr) linkage_map<-linkage_map[,3] C1<-(No_mrk_in_each_chr[1]+No_mrk_in_each_chr[2])*2 C2<-sum(No_mrk_in_each_chr)*2 mat<-rp_3$output[[generation+1]]$mrk mat<-mat[,-c(1,2)] mat<-mat[,(c1+1):c2] # Mean r2 rld<-calc_ld(mat=mat,maf=0.1,method='adjacent') # LD decay rld<-calc_ld(mat=mat,maf=0.1,method='pairwise', LD_summary=TRUE,linkage_map=linkage_map,interval=1) rld$ld_decay Example 5: Two-way crossbreeding Simulation of a two-way crossbreeding program. The crossbreeding scheme in this example involves three steps: Step 1: Historical population is created. Step 2: Two recent populations named as Breed A and B are created by sampling individuals from historical population. Step 3: Breed A and B are crossed. Schematic representation of the two-way crossbreeding in this example is in Figure 4. # # STEP 1: CREATE HISTORICAL POPULATION # Genome consisted of 4 chromosomes 14
15 genome<-data.frame(matrix(na, nrow=4, ncol=6)) names(genome)<-c("chr","len","nmrk","mpos","nqtl","qpos") genome$chr<-c(1:4) genome$len<-c(94,110,111.5,78.3) genome$nmrk<-c(500,500,500,500) genome$mpos<-rep('rnd',4) genome$nqtl<-c(70,80,90,65) genome$qpos<-rep('rnd',4) genome historical<-make_hp(hpsize=200,ng=300,h2=0.25,d2=0.10,phen_var=1,genome=genome,mutr=5*10**-4,sel_seq_qtl=0.1,sel_seq_mrk=0.05,laf=0.5) # # STEP 2: MAKE BREED A AND B # BREED A Breed_A_Male_fndrs<-data.frame(number=50,select='rnd') Breed_A_Female_fndrs<-data.frame(number=50,select='rnd') # Selection and matings in Breed A # Selection of 50 sires and 100 dam # Selection criteria is "rnd" for both sires and dams Selection<-data.frame(matrix(NA, nrow=2, ncol=2)) names(selection)<-c('number','type') Selection$Number[1:2]<-c(50,100) Selection$type[1:2]<-c('rnd','rnd') Selection # Save information for the last generation of Breed A Breed_A_data<-data.frame(matrix(NA, nrow=1, ncol=6)) names(breed_a_data)<-c('data','qtl','marker','seq','freq_qtl','freq_mrk') Breed_A_data[,1]<-10 Breed_A_data[,2]<-10 Breed_A_data[,3]<-10 Breed_A_data[,4]<-10 Breed_A_data[,5]<-10 Breed_A_data[,6]<-10 Breed_A_data Breed_A<-sample_hp(hp_out=historical,Male_founders= Breed_A_Male_fndrs,Female_founders=Breed_A_Female_fndrs, ng=10,selection=selection, litter_size=5,saveat="breeda",sh_output=breed_a_data,display=true) # BREED B Breed_B_Male_fndrs<-data.frame(number=50,select='rnd') Breed_B_Female_fndrs<-data.frame(number=50,select='rnd') 15
16 # Selection and matings in Breed B # Selection of 50 sires and 100 dam # Selection criteria is "phen" for both sires and dams Selection<-data.frame(matrix(NA, nrow=2, ncol=3)) names(selection)<-c('number','type','value') Selection$Number[1:2]<-c(50,100) Selection$type[1:2]<-c('phen','phen') Selection$Value[1:2]<-c('h','h') Selection Breed_B<-sample_hp(hp_out=historical,Male_founders= Breed_B_Male_fndrs,Female_founders=Breed_B_Female_fndrs, ng=10,selection=selection, litter_size=5) # # STEP 3: CROSSING BETWEEN BREED A AND B # Selection of founders in crossbreeding for Breed A # Selection of 100 sires/dams from last generation of Breed_A in step 2. # Selection criteria is "rnd" for both sires and dams founder_pop1<-data.frame(matrix(na, nrow=2, ncol=3)) names(founder_pop1)<-c('size','generation','select') founder_pop1[1,]<-c(100,10,'rnd') founder_pop1[2,]<-c(100,10,'rnd') founder_pop1 # Selection of founders in crossbreeding for Breed B # Selection of 80 sires and 150 dams # Selection criteria is "phen" for sires # Selection criteria is "rnd" for dams founder_pop2<-data.frame(matrix(na, nrow=2, ncol=4)) names(founder_pop2)<-c('size','generation','select','value') founder_pop2[1,]<-c(80,10,'phen','h') founder_pop2[2,]<-c(150,10,'rnd','h') # "h" will be ignored as SC is "rnd" founder_pop2 # Selection of animals from founder_pop1 and founder_pop2 to be crossed founder_cross<-data.frame(matrix(na, nrow=2, ncol=4)) names(founder_cross)<-c('pop','size','select','value') founder_cross[1,]<-c('pop1',50,'tbv','h') # Select males from Breed A founder_cross[2,]<-c('pop2',100,'phen','h') # Select females from Breed B founder_cross # Selection scheme in Breed A to produce purebred replacement animals Selection_pop1<-data.frame(matrix(NA, nrow=2, ncol=3)) 16
17 names(selection_pop1)<-c('number','type','value') Selection_pop1$Number[1:2]<-c(100,100) Selection_pop1$type[1:2]<-c('gebv','gebv') Selection_pop1$Value[1:2]<-c('h','h') Selection_pop1 # Selection scheme in Breed B to produce purebred replacement animals Selection_pop2<-data.frame(matrix(NA, nrow=2, ncol=3)) names(selection_pop2)<-c('number','type','value') Selection_pop2$Number[1:2]<-c(100,100) Selection_pop2$type[1:2]<-c('gebv','gebv') Selection_pop2$Value[1:2]<-c('h','h') Selection_pop2 # Selection scheme for crossing between A and B Cross_design<-data.frame(matrix(NA, nrow=2, ncol=4)) names(cross_design)<-c('pop','size','select','value') Cross_design[1,]<-c('pop1',100,'gebvc','h') Cross_design[2,]<-c('pop2',200,'gebvc','h') Cross_design # Training in Breed A train_a<-data.frame(matrix(na, nrow=1, ncol=5)) names(train_a)<-c('size','sel','method','niter','show') train_a$size<-500 train_a$sel<-'rnd' train_a$method<-'brr' train_a$niter<-1000 train_a$show<-true train_a # Training in Breed B train_b<-data.frame(matrix(na, nrow=1, ncol=5)) names(train_b)<-c('size','sel','method','niter','show') train_b$size<-500 train_b$sel<-'rnd' train_b$method<-'brr' train_b$niter<-1000 train_b$show<-true train_b # Save information for crossbred AB for the last 2 generations output_cross<-data.frame(matrix(na, nrow=4, ncol=5)) names(output_cross)<-c('data','qtl','marker','freq_qtl','freq_mrk') output_cross[,1]<-c(6,7) output_cross[,2]<-c(6,7) output_cross[,3]<-c(6,7) 17
18 output_cross[,4]<-c(6,7) output_cross[,5]<-c(6,7) output_cross cross_ab<-xbreed(pop1=breed_a,pop2=breed_b,founder_pop1= founder_pop1,founder_pop2=founder_pop2, founder_cross=founder_cross, Selection_pop1=Selection_pop1,Selection_pop2=Selection_pop2, Cross_design=Cross_design,train_type='purebred', train_pop1=train_a,train_pop2=train_b,ng=7,litter_size=5, saveat='cross_pop',output_cross=output_cross,display=true) # SOME OUTPUT cross_ab$summary_data_pop1 # Summary Breed A cross_ab$summary_data_pop1 # Summary Breed B cross_ab$summary_data_cross # Summary of AB crossbred # Data for the last generation of BREED A cross_ab$pop1[[8]]$data # Data for the last generation of BREED B cross_ab$pop2[[8]]$data # Data for the last generation of crossbreds cross_ab$cross$output[[8]]$data Example 6: Three-way crossbreeding Simulation of a three-way crossbreeding program. The crossbreeding scheme in this example involves four steps: Step 1: Historical population is created. Step 2: Three recent populations named as Breed A, B and C are created by sampling individuals from last generation of historical population. Step 3: Breed A and B are crossed. Step 4: Breed C as sire line is crossed to the female crossbred progeny of Breed A and B named as (AB). # # STEP 1: CREATE HISTORICAL POPULATION # Genome consisted of 2 chromosomes genome<-data.frame(matrix(na, nrow=2, ncol=6)) names(genome)<-c("chr","len","nmrk","mpos","nqtl","qpos") genome$chr<-c(1:2) genome$len<-c(94,110) genome$nmrk<-c(1100,1050) genome$mpos<-c('even','rnd') genome$nqtl<-c(90,100) 18
19 Figure 4: Schematic representation of the two-way crossbreeding. 19
20 genome$qpos<-c('even','rnd') genome historical<-make_hp(hpsize=300,ng=400,h2=0.5,d2=0.10,phen_var=1,genome=genome,mutr=5*10**-4,sel_seq_qtl=0.1,sel_seq_mrk=0.05,laf=0.5) # # STEP 2: MAKE BREED A, B AND C # BREED A Breed_A_Male_fndrs<-data.frame(number=50,select='rnd') Breed_A_Female_fndrs<-data.frame(number=50,select='rnd') # Selection scheme in Breed A Selection<-data.frame(matrix(NA, nrow=2, ncol=2)) names(selection)<-c('number','type') Selection$Number[1:2]<-c(50,100) Selection$type[1:2]<-c('rnd','rnd') Selection Breed_A<-sample_hp(hp_out=historical,Male_founders= Breed_A_Male_fndrs,Female_founders=Breed_A_Female_fndrs, ng=10,selection=selection, litter_size=4,display=true) # BREED B Breed_B_Male_fndrs<-data.frame(number=50,select='rnd') Breed_B_Female_fndrs<-data.frame(number=50,select='rnd') # Selection scheme in Breed B Selection<-data.frame(matrix(NA, nrow=2, ncol=3)) names(selection)<-c('number','type','value') Selection$Number[1:2]<-c(50,50) Selection$type[1:2]<-c('phen','phen') Selection$Value[1:2]<-c('h','h') Selection Breed_B<-sample_hp(hp_out=historical,Male_founders= Breed_B_Male_fndrs,Female_founders=Breed_B_Female_fndrs, ng=10,selection=selection, litter_size=5) # BREED C Breed_C_Male_fndrs<-data.frame(number=50,select='rnd') Breed_C_Female_fndrs<-data.frame(number=50,select='rnd') # Selection scheme in Breed C Selection<-data.frame(matrix(NA, nrow=2, ncol=3)) 20
21 names(selection)<-c('number','type','value') Selection$Number[1:2]<-c(50,50) Selection$type[1:2]<-c('tbv','tbv') Selection$Value[1:2]<-c('h','h') Selection Breed_C<-sample_hp(hp_out=historical,Male_founders= Breed_C_Male_fndrs,Female_founders=Breed_C_Female_fndrs, ng=5,selection=selection, litter_size=5) # # STEP 3: CROSSING BETWEEN BREED A AND B # Selection of founders in crossbreeding for Breed A # Selection of 100 sires and 100 dams # Selection criteria is "phen" for both sires and dams founder_pop1<-data.frame(matrix(na, nrow=2, ncol=4)) names(founder_pop1)<-c('size','generation','select','value') founder_pop1[1,]<-c(100,10,'phen','h') founder_pop1[2,]<-c(100,10,'phen','h') founder_pop1 # Selection of founders in crossbreeding for Breed B # Selection of 80 sires and 100 dams # Selection criteria is "rnd" for both sires and dams founder_pop2<-data.frame(matrix(na, nrow=2, ncol=3)) names(founder_pop2)<-c('size','generation','select') founder_pop2[1,]<-c(80,10,'rnd') founder_pop2[2,]<-c(100,10,'rnd') founder_pop2 # Selection of animals from founder_pop1 and founder_pop2 to be crossed founder_cross<-data.frame(matrix(na, nrow=2, ncol=4)) names(founder_cross)<-c('pop','size','select','value') founder_cross[1,]<-c('pop1',50,'rnd','h') # Select males from Breed A founder_cross[2,]<-c('pop2',100,'phen','h') # Select females from Breed B founder_cross # Selection scheme in Breed A to produce purebred replacement animals Selection_pop1<-data.frame(matrix(NA, nrow=2, ncol=3)) names(selection_pop1)<-c('number','type','value') Selection_pop1$Number[1:2]<-c(100,100) Selection_pop1$type[1:2]<-c('gebv','gebv') Selection_pop1$Value[1:2]<-c('h','h') Selection_pop1 # Selection scheme in Breed B to produce purebred replacement animals 21
22 Selection_pop2<-data.frame(matrix(NA, nrow=2, ncol=3)) names(selection_pop2)<-c('number','type','value') Selection_pop2$Number[1:2]<-c(100,100) Selection_pop2$type[1:2]<-c('gebv','gebv') Selection_pop2$Value[1:2]<-c('h','h') Selection_pop2 # Selection scheme for crossing between A and B Cross_design<-data.frame(matrix(NA, nrow=2, ncol=4)) names(cross_design)<-c('pop','size','select','value') Cross_design[1,]<-c('pop1',100,'gebvc','h') Cross_design[2,]<-c('pop2',100,'gebvc','h') Cross_design # Training in Breed A train_a<-data.frame(matrix(na, nrow=1, ncol=5)) names(train_a)<-c('size','sel','method','niter','show') train_a$size<-500 train_a$sel<-'rnd' train_a$method<-'brr' train_a$niter<-1000 train_a$show<-true train_a # Training in Breed B train_b<-data.frame(matrix(na, nrow=1, ncol=5)) names(train_b)<-c('size','sel','method','niter','show') train_b$size<-500 train_b$sel<-'rnd' train_b$method<-'brr' train_b$niter<-1000 train_b$show<-true train_b # Save information for crossbred AB output_cross<-data.frame(matrix(na, nrow=4, ncol=5)) names(output_cross)<-c('data','qtl','marker','freq_qtl','freq_mrk') output_cross[,1]<-c(1,2,6,7) output_cross[,2]<-c(1,2,6,7) output_cross[,3]<-c(1,2,6,7) output_cross[,4]<-c(1,2,6,7) output_cross[,5]<-c(1,2,6,7) output_cross cross_ab<-xbreed(pop1=breed_a,pop2=breed_b,founder_pop1= founder_pop1,founder_pop2=founder_pop2, founder_cross=founder_cross, 22
23 Selection_pop1=Selection_pop1,Selection_pop2=Selection_pop2, Cross_design=Cross_design,train_type='purebred', train_pop1=train_a,train_pop2=train_b,ng=7,litter_size=5, saveat='cross_pop',output_cross=output_cross,display=true) # # STEP 4: CROSSING BETWEEN AB CROSSBREDS AND BREED C # Selection of founders from AB crossbreds # Selection of 100 sires and 100 dams # Selection criteria is "rnd" for sires and "tbv" for dams founder_pop1<-data.frame(matrix(na, nrow=2, ncol=4)) names(founder_pop1)<-c('size','generation','select','value') founder_pop1[1,]<-c(100,7,'rnd','l') # "l" will be ignored as SC is "rnd" founder_pop1[2,]<-c(100,7,'tbv','h') founder_pop1 # Selection of founders from Breed C # Selection of 80 sires and 100 dams # Selection criteria is "phen" for sires # Selection criteria is "rnd" dams founder_pop2<-data.frame(matrix(na, nrow=2, ncol=4)) names(founder_pop2)<-c('size','generation','select','value') founder_pop2[1,]<-c(80,5,'phen','h') founder_pop2[2,]<-c(100,5,'rnd','l') founder_pop2 # # Selection of animals from founder_pop1 (crossbred AB) # and founder_pop2 (Breed C) to be crossed founder_cross<-data.frame(matrix(na, nrow=2, ncol=4)) names(founder_cross)<-c('pop','size','select','value') founder_cross[1,]<-c('pop1',50,'rnd','h') # Select males from Breed C founder_cross[2,]<-c('pop2',100,'phen','h') # Select females from cross of AB founder_cross # Selection scheme in AB crossbreds for subsequent generations Selection_pop1<-data.frame(matrix(NA, nrow=2, ncol=3)) names(selection_pop1)<-c('number','type','value') Selection_pop1$Number[1:2]<-c(100,100) Selection_pop1$type[1:2]<-c('gebv','gebv') Selection_pop1$Value[1:2]<-c('h','h') Selection_pop1 # Selection scheme in Breed C to produce purebred replacement animals Selection_pop2<-data.frame(matrix(NA, nrow=2, ncol=3)) names(selection_pop2)<-c('number','type','value') Selection_pop2$Number[1:2]<-c(100,100) Selection_pop2$type[1:2]<-c('gebv','gebv') 23
24 Selection_pop2$Value[1:2]<-c('h','h') Selection_pop2 # Selection scheme for crossing between AB and C Cross_design<-data.frame(matrix(NA, nrow=2, ncol=4)) names(cross_design)<-c('pop','size','select','value') Cross_design[1,]<-c('pop2',50,'gebvc','h') # Select males from Breed C Cross_design[2,]<-c('pop1',100,'gebvc','h') # Select females from (AB) Cross_design # Training in Crossbred AB(C) train<-data.frame(matrix(na, nrow=1, ncol=5)) names(train)<-c('size','sel','method','niter','show') train$size<-500 train$sel<-'rnd' train$method<-'bl' train$niter<-1000 train$show<-false # Iteration history won't be displayed train # Save information for crossbred (AB)C output_cross<-data.frame(matrix(na, nrow=3, ncol=4)) names(output_cross)<-c('data','qtl','freq_qtl','freq_mrk') output_cross[,1]<-c(1,2,3) output_cross[,2]<-c(1,2,3) output_cross[,3]<-c(1,2,3) output_cross[,4]<-c(1,2,3) output_cross cross_ab_c<-xbreed(pop1=cross_ab$cross,pop2=breed_c,founder_pop1= founder_pop1,founder_pop2=founder_pop2, founder_cross=founder_cross, Selection_pop1=Selection_pop1,Selection_pop2=Selection_pop2, Cross_design=Cross_design,train_type='crossbred', train_cross=train,ng=3,litter_size=5, saveat='cross_(ab)_c',output_cross=output_cross) # SOME OUTPUT cross_ab_c$summary_data_pop1 # Summary Breed AB cross_ab_c$summary_data_pop1 # Summary Breed C cross_ab_c$summary_data_cross # Summary of ABC crossbred # Data for the last generation of crossbreds ABC cross_ab_c$cross$output[[4]]$data 24
25 Figure 5: Schematic representation of the three-way crossbreeding. 25
26 Example 7: Four-way crossbreeding Simulation of four-way crossbreeding program for 8 generations. The crossbreeding scheme in this example involves five steps: Step 1: Historical population is created. Step 2: Four recent populations named as Breed A, B, C and D are created by sampling individuals from the last generation of historical population. Step 3: Breed A and B are crossed. Step 4: Breed C and D are crossed. Step 5: Crossbred males of AB are crossed to crossbred females of CD. # # STEP 1: CREATE HISTORICAL POPULATION # Genome consisted of 2 chromosomes genome<-data.frame(matrix(na, nrow=2, ncol=6)) names(genome)<-c("chr","len","nmrk","mpos","nqtl","qpos") genome$chr<-c(1:2) genome$len<-c(94,110) genome$nmrk<-c(800,800) genome$mpos<-c('rnd','rnd') genome$nqtl<-c(100,100) genome$qpos<-c('rnd','rnd') genome historical<-make_hp(hpsize=400,ng=200,h2=0.5,d2=0.10,phen_var=1,genome=genome,mutr=5*10**-4,sel_seq_qtl=0.1,sel_seq_mrk=0.05,laf=0.5) # # STEP 2: MAKE BREED A, B, C AND D # BREED A Breed_A_Male_fndrs<-data.frame(number=50,select='rnd') Breed_A_Female_fndrs<-data.frame(number=50,select='rnd') # Selection and matings in Breed A Selection<-data.frame(matrix(NA, nrow=2, ncol=2)) names(selection)<-c('number','type') Selection$Number[1:2]<-c(50,50) Selection$type[1:2]<-c('rnd','rnd') Selection Breed_A<-sample_hp(hp_out=historical,Male_founders= Breed_A_Male_fndrs,Female_founders=Breed_A_Female_fndrs, ng=5,selection=selection,litter_size=5) # BREED B Breed_B_Male_fndrs<-data.frame(number=50,select='rnd') Breed_B_Female_fndrs<-data.frame(number=50,select='rnd') 26
27 # Selection and matings in Breed B Selection<-data.frame(matrix(NA, nrow=2, ncol=2)) names(selection)<-c('number','type') Selection$Number[1:2]<-c(50,50) Selection$type[1:2]<-c('rnd','rnd') Selection Breed_B<-sample_hp(hp_out=historical,Male_founders= Breed_B_Male_fndrs,Female_founders=Breed_B_Female_fndrs, ng=5,selection=selection,litter_size=5) # BREED C Breed_C_Male_fndrs<-data.frame(number=50,select='rnd') Breed_C_Female_fndrs<-data.frame(number=50,select='rnd') # Selection and matings in Breed C Selection<-data.frame(matrix(NA, nrow=2, ncol=2)) names(selection)<-c('number','type') Selection$Number[1:2]<-c(50,50) Selection$type[1:2]<-c('rnd','rnd') Selection Breed_C<-sample_hp(hp_out=historical,Male_founders= Breed_C_Male_fndrs,Female_founders=Breed_C_Female_fndrs, ng=5,selection=selection,litter_size=5) # BREED D Breed_D_Male_fndrs<-data.frame(number=50,select='rnd') Breed_D_Female_fndrs<-data.frame(number=50,select='rnd') # Selection and matings in Breed D Selection<-data.frame(matrix(NA, nrow=2, ncol=2)) names(selection)<-c('number','type') Selection$Number[1:2]<-c(50,50) Selection$type[1:2]<-c('rnd','rnd') Selection Breed_D<-sample_hp(hp_out=historical,Male_founders= Breed_D_Male_fndrs,Female_founders=Breed_D_Female_fndrs, ng=5,selection=selection,litter_size=5) # # STEP 3: CROSSING BETWEEN BREED A AND B # Selection of founders in crossbreeding for Breed A # Selection of 25 sires and 50 dams # Selection criteria is "rnd" for both sires and dams 27
28 founder_pop1<-data.frame(matrix(na, nrow=2, ncol=4)) names(founder_pop1)<-c('size','generation','select','value') founder_pop1[1,]<-c(25,5,'phen','h') founder_pop1[2,]<-c(50,5,'phen','h') founder_pop1 # Selection of founders in crossbreeding for Breed B # Selection of 25 sires and 50 dams # Selection criteria is "rnd" for sires and dams # Selection criteria is "rnd" dams founder_pop2<-data.frame(matrix(na, nrow=2, ncol=3)) names(founder_pop2)<-c('size','generation','select') founder_pop2[1,]<-c(25,5,'rnd') founder_pop2[2,]<-c(50,5,'rnd') founder_pop2 # # Selection of animals from founder_pop1 (Breed A) # and founder_pop2 (Breed B) to be crossed founder_cross<-data.frame(matrix(na, nrow=2, ncol=4)) names(founder_cross)<-c('pop','size','select','value') founder_cross[1,]<-c('pop1',25,'tbv','h') # Select males from Breed A founder_cross[2,]<-c('pop2',50,'tbv','h') # Select females from Breed B founder_cross # Selection scheme in Breed A to produce purebred replacement animals Selection_pop1<-data.frame(matrix(NA, nrow=2, ncol=3)) names(selection_pop1)<-c('number','type','value') Selection_pop1$Number[1:2]<-c(50,50) Selection_pop1$type[1:2]<-c('tbv','phen') Selection_pop1$Value[1:2]<-c('h','h') Selection_pop1 # Selection scheme in Breed B to produce purebred replacement animals Selection_pop2<-data.frame(matrix(NA, nrow=2, ncol=3)) names(selection_pop2)<-c('number','type','value') Selection_pop2$Number[1:2]<-c(50,50) Selection_pop2$type[1:2]<-c('phen','tbv') Selection_pop2$Value[1:2]<-c('h','h') Selection_pop2 # Selection scheme for crossing between A and B Cross_design<-data.frame(matrix(NA, nrow=2, ncol=4)) names(cross_design)<-c('pop','size','select','value') Cross_design[1,]<-c('pop1',10,'phen','h') Cross_design[2,]<-c('pop2',100,'phen','h') Cross_design 28
29 cross_ab<-xbreed(pop1=breed_a,pop2=breed_b,founder_pop1= founder_pop1,founder_pop2=founder_pop2, founder_cross=founder_cross, Selection_pop1=Selection_pop1,Selection_pop2=Selection_pop2, Cross_design=Cross_design,ng=7,litter_size=4) # # STEP 4: CROSSING BETWEEN BREED C AND D # Selection of founders in crossbreeding for Breed C # Selection of 100 sires and 100 dams # Selection criteria is "rnd" for both sires and dams founder_pop1<-data.frame(matrix(na, nrow=2, ncol=4)) names(founder_pop1)<-c('size','generation','select','value') founder_pop1[1,]<-c(25,5,'phen','h') founder_pop1[2,]<-c(50,5,'phen','h') founder_pop1 # Selection of founders in crossbreeding for Breed D # Selection of 45 sires and 80 dams # Selection criteria is "rnd" for both sires and dams founder_pop2<-data.frame(matrix(na, nrow=2, ncol=3)) names(founder_pop2)<-c('size','generation','select') founder_pop2[1,]<-c(25,5,'rnd') founder_pop2[2,]<-c(50,5,'rnd') founder_pop2 # Selection of animals from founder_pop1 and founder_pop2 to be crossed founder_cross<-data.frame(matrix(na, nrow=2, ncol=4)) names(founder_cross)<-c('pop','size','select','value') founder_cross[1,]<-c('pop1',25,'tbv','h') # Select males from Breed C founder_cross[2,]<-c('pop2',50,'tbv','h') # Select females from Breed D founder_cross # Selection scheme in Breed C to produce purebred replacement animals Selection_pop1<-data.frame(matrix(NA, nrow=2, ncol=3)) names(selection_pop1)<-c('number','type','value') Selection_pop1$Number[1:2]<-c(50,50) Selection_pop1$type[1:2]<-c('tbv','phen') Selection_pop1$Value[1:2]<-c('h','h') Selection_pop1 # Selection scheme in Breed D to produce purebred replacement animals Selection_pop2<-data.frame(matrix(NA, nrow=2, ncol=3)) names(selection_pop2)<-c('number','type','value') Selection_pop2$Number[1:2]<-c(50,50) Selection_pop2$type[1:2]<-c('phen','tbv') Selection_pop2$Value[1:2]<-c('h','h') 29
30 Selection_pop2 # Selection scheme for crossing between C and D Cross_design<-data.frame(matrix(NA, nrow=2, ncol=4)) names(cross_design)<-c('pop','size','select','value') Cross_design[1,]<-c('pop1',10,'phen','h') Cross_design[2,]<-c('pop2',100,'phen','h') Cross_design cross_cd<-xbreed(pop1=breed_c,pop2=breed_d,founder_pop1= founder_pop1,founder_pop2=founder_pop2, founder_cross=founder_cross, Selection_pop1=Selection_pop1,Selection_pop2=Selection_pop2, Cross_design=Cross_design,ng=7,litter_size=5) # SOME OUTPUT cross_cd$summary_data_pop1 # Summary Breed C cross_cd$summary_data_pop1 # Summary Breed D cross_cd$summary_data_cross # Summary of CD crossbred # Data for the CD crossbreds cross_cd$pop1[[2]]$data # generation 1 cross_cd$pop1[[5]]$data # generation 4 # # STEP 5: CROSSING MALES OF (AB) TO FEMALES OF (CD) # Selection of founders from AB # Selection of 50 sires and 50 dams # Selection criteria is "phen" for both sires and dams founder_pop1<-data.frame(matrix(na, nrow=2, ncol=4)) names(founder_pop1)<-c('size','generation','select','value') founder_pop1[1,]<-c(50,7,'phen','h') founder_pop1[2,]<-c(50,7,'phen','l') founder_pop1 # Selection of founders from CD # Selection of 40 sires and 100 dams # Selection criteria is "gebv" for sires and dams # Selection criteria is "rnd" dams founder_pop2<-data.frame(matrix(na, nrow=2, ncol=4)) names(founder_pop2)<-c('size','generation','select','value') founder_pop2[1,]<-c(40,7,'tbv','h') founder_pop2[2,]<-c(80,7,'tbv','l') founder_pop2 # Selection of animals from founder_pop1 and founder_pop2 to be crossed 30
31 founder_cross<-data.frame(matrix(na, nrow=2, ncol=4)) names(founder_cross)<-c('pop','size','select','value') founder_cross[1,]<-c('pop1',50,'rnd','h') # Select males from Breed C founder_cross[2,]<-c('pop2',100,'phen','h') # Select females from cross of AB founder_cross # Selection scheme in AB Selection_pop1<-data.frame(matrix(NA, nrow=2, ncol=3)) names(selection_pop1)<-c('number','type','value') Selection_pop1$Number[1:2]<-c(100,100) Selection_pop1$type[1:2]<-c('phen','phen') Selection_pop1$Value[1:2]<-c('h','h') Selection_pop1 # Selection scheme in CD Selection_pop2<-data.frame(matrix(NA, nrow=2, ncol=3)) names(selection_pop2)<-c('number','type','value') Selection_pop2$Number[1:2]<-c(100,100) Selection_pop2$type[1:2]<-c('phen','phen') Selection_pop2$Value[1:2]<-c('h','h') Selection_pop2 # Selection scheme for crossing between AB and CD Cross_design<-data.frame(matrix(NA, nrow=2, ncol=4)) names(cross_design)<-c('pop','size','select','value') Cross_design[1,]<-c('pop1',50,'tbv','h') Cross_design[2,]<-c('pop2',100,'tbv','h') Cross_design # Save information for crossbred ABCD output_cross<-data.frame(matrix(na, nrow=4, ncol=5)) names(output_cross)<-c('data','qtl','marker','freq_qtl','freq_mrk') output_cross[,1]<-c(1) output_cross[,2]<-c(1) output_cross[,3]<-c(1) output_cross[,4]<-c(1) output_cross[,5]<-c(1) output_cross cross_ab_cd<-xbreed(pop1=cross_ab$cross,pop2=cross_cd$cross,founder_pop1= founder_pop1,founder_pop2=founder_pop2, founder_cross=founder_cross, Selection_pop1=Selection_pop1,Selection_pop2=Selection_pop2, Cross_design=Cross_design,ng=1,litter_size=5, saveat='cross_(ab)_(cd)',output_cross=output_cross) 31
32 # SOME OUTPUT cross_ab_cd$summary_data_pop1 # Summary AB crossbreds cross_ab_cd$summary_data_pop1 # Summary CD crossbreds cross_ab_cd$summary_data_cross # Summary of ABCD crossbred # Data for the ABCD crossbreds cross_ab_cd$pop1[[1]]$data cross_ab_cd$pop1[[2]]$data Figure 6: Schematic representation of the four-way crossbreeding. Acknowledgments Development of xbreed was supported by the Danish Strategic Research Council (GenSAP: Centre for Genomic Selection in Animals and Plants, contract no ). 32
33 References Crow, J.F., and M. Kimura, An introduction to population genetics theory. Caldwell, NJ: The Blackburn Press Hill, W.G., and A. Robertson, Linkage disequilibrium in finite populations. Theor. Appl. Genet. 38: Hudson, R., Generating samples under a Wright-Fisher neutral model. Bioinformatics 18: Tenesa, A., P. Navarro, B. J. Hayes, D. L. Duffy, G. M. Clarke et al., Recent human effective population size estimated from linkage disequilibrium. Genome Res. 17: Sved, J. A., Linkage disequilibrium and homozygosity of chromosome segments in finite populations. Theor. Popul. Biol. 2: Wright, S., The distribution of gene frequencies in populations. Genetics 23,
Assigning breed origin to alleles in crossbred animals
 DOI 10.1186/s12711-016-0240-y Genetics Selection Evolution RESEARCH ARTICLE Open Access Assigning breed origin to alleles in crossbred animals Jérémie Vandenplas 1*, Mario P. L. Calus 1, Claudia A. Sevillano
DOI 10.1186/s12711-016-0240-y Genetics Selection Evolution RESEARCH ARTICLE Open Access Assigning breed origin to alleles in crossbred animals Jérémie Vandenplas 1*, Mario P. L. Calus 1, Claudia A. Sevillano
7.013 Problem Set
 7.013 Problem Set 2-2013 Question 1 You are working with the following parental flies (P1-P4). P1: Light body color and normal wings P2: Light body color and wingless P3: Dark body color and wingless P4:
7.013 Problem Set 2-2013 Question 1 You are working with the following parental flies (P1-P4). P1: Light body color and normal wings P2: Light body color and wingless P3: Dark body color and wingless P4:
First results on genomic selection in French show-jumping horses
 EAAP 2012 First results on genomic selection in French show-jumping horses A. Ricard 1,3, S. Danvy 3, A. Legarra 2 1 Biologie Intégrative et Génétique Equine, UMR 1313 GABI, INRA 2 Station d amélioration
EAAP 2012 First results on genomic selection in French show-jumping horses A. Ricard 1,3, S. Danvy 3, A. Legarra 2 1 Biologie Intégrative et Génétique Equine, UMR 1313 GABI, INRA 2 Station d amélioration
Across-breed genomic evaluation based on BovineHD genotypes, and phenotypes of bulls and cows
 Across-breed genomic evaluation based on BovineHD genotypes, and phenotypes of bulls and cows Chris Schrooten 1), Ghyslaine Schopen 1), Phil Beatson 2) 1) CRV BV, The Netherlands 2) CRV Ambreed, New Zealand
Across-breed genomic evaluation based on BovineHD genotypes, and phenotypes of bulls and cows Chris Schrooten 1), Ghyslaine Schopen 1), Phil Beatson 2) 1) CRV BV, The Netherlands 2) CRV Ambreed, New Zealand
Challenges related to introducing genomic selection in sport horse breeding EAAP 2015
 Challenges related to introducing genomic selection in sport horse breeding EAAP 2015 Anne Ricard, INRA, UMR 1313, IFCE Recherche et Innovation, anne.ricard@toulouse.inra.fr «That is the question» Genomic
Challenges related to introducing genomic selection in sport horse breeding EAAP 2015 Anne Ricard, INRA, UMR 1313, IFCE Recherche et Innovation, anne.ricard@toulouse.inra.fr «That is the question» Genomic
Understanding Genetics
 Understanding Genetics INDEX Understanding genetics 3 Carrier mating disease risks 5 While this booklet focuses on cattle as an example, this information is useful for all species. DNA is essential to
Understanding Genetics INDEX Understanding genetics 3 Carrier mating disease risks 5 While this booklet focuses on cattle as an example, this information is useful for all species. DNA is essential to
Genetics Discussion for the American Black Hereford Association. David Greg Riley Texas A&M University
 Genetics Discussion for the American Black Hereford Association David Greg Riley Texas A&M University Topics Managing Genetic Defects in Cattle Populations Contemporary Groupings Data Collection and Reporting
Genetics Discussion for the American Black Hereford Association David Greg Riley Texas A&M University Topics Managing Genetic Defects in Cattle Populations Contemporary Groupings Data Collection and Reporting
DNA Breed Profile Testing FAQ. What is the history behind breed composition testing?
 DNA Breed Profile Testing FAQ What is the history behind breed composition testing? Breed composition testing, or breed integrity testing has been in existence for many years. Hampshire, Landrace, and
DNA Breed Profile Testing FAQ What is the history behind breed composition testing? Breed composition testing, or breed integrity testing has been in existence for many years. Hampshire, Landrace, and
Do not write in margin QUESTIONSHEET 1
 QUESTIONSHEET 1 Mendel s first law states that only one of a pair of contrasting characters may be represented in a single gamete. His second law states that either of a pair of contracting characters
QUESTIONSHEET 1 Mendel s first law states that only one of a pair of contrasting characters may be represented in a single gamete. His second law states that either of a pair of contracting characters
Introduction to be read or described to the participants:
 Introduction to be read or described to the participants: When breeding animals, it is important to understand the reasons why the offspring (foal, lamb, calf, etc.) look a certain way or have the traits
Introduction to be read or described to the participants: When breeding animals, it is important to understand the reasons why the offspring (foal, lamb, calf, etc.) look a certain way or have the traits
Dehorning cattle via genetics
 IRISH CATTLE BREEDING FEDERATION Dehorning cattle via genetics Tuesday 11 th April Portlaoise 1 Outline Background Genetics of Polled gene Polled vs Horned How to increase Polled gene Breeding for the
IRISH CATTLE BREEDING FEDERATION Dehorning cattle via genetics Tuesday 11 th April Portlaoise 1 Outline Background Genetics of Polled gene Polled vs Horned How to increase Polled gene Breeding for the
Figure S1a Diagram of the HA412- HO x RHA415 cross. J. E. Bowers et al.
 Figure S1a Diagram of the HA412- HO x RHA415 cross 1 SI Figure S1b Diagram of the HA412- HO x RHA415 cross 2 SI Figure S1c Diagram of the RHA280 x RHA801 cross 3 SI Figure S1d Diagram of the NMS373 x Hopi
Figure S1a Diagram of the HA412- HO x RHA415 cross 1 SI Figure S1b Diagram of the HA412- HO x RHA415 cross 2 SI Figure S1c Diagram of the RHA280 x RHA801 cross 3 SI Figure S1d Diagram of the NMS373 x Hopi
Genetic & Breeding Concepts Sheep and Goat Expo 2017 San Angelo
 Presentation Overview Genetic & Breeding Concepts Sheep and Goat Expo 2017 San Angelo Andy D. Herring Department of Animal Science andy.herring@tamu.com General breeding and genetics terms and concepts
Presentation Overview Genetic & Breeding Concepts Sheep and Goat Expo 2017 San Angelo Andy D. Herring Department of Animal Science andy.herring@tamu.com General breeding and genetics terms and concepts
Estimating breeding value
 Estimating breeding value Key terms Estimated breeding value (EBV) Heritability Contemporary groups Using information from relatives Accuracy Which animal to select? Animal P1 P2 EBV sire's perform. Index
Estimating breeding value Key terms Estimated breeding value (EBV) Heritability Contemporary groups Using information from relatives Accuracy Which animal to select? Animal P1 P2 EBV sire's perform. Index
Implementing Genomic Information in Breeding Schemes of Danish Warmblood Horses
 Implementing Genomic Information in Breeding Schemes of Danish Warmblood Horses MSc Thesis in Agrobiology 45 ECTS Faculty of Science and Technology, Department of Molecular Biology and Genetics - Centre
Implementing Genomic Information in Breeding Schemes of Danish Warmblood Horses MSc Thesis in Agrobiology 45 ECTS Faculty of Science and Technology, Department of Molecular Biology and Genetics - Centre
Genetic analyses of the Franches-Montagnes horse breed with genome-wide SNP data
 Photo: M. Rindlisbacher Genetic analyses of the Franches-Montagnes horse breed with genome-wide SNP data H. Hasler 1, C. Flury 1, B. Haase 2, D. Burger 3, C.Stricker 4, H. Simianer 5, T. Leeb 2, S. Rieder
Photo: M. Rindlisbacher Genetic analyses of the Franches-Montagnes horse breed with genome-wide SNP data H. Hasler 1, C. Flury 1, B. Haase 2, D. Burger 3, C.Stricker 4, H. Simianer 5, T. Leeb 2, S. Rieder
X-Sheet 4 Genetics: Inheritance and Terminology
 X-Sheet 4 Genetics: Inheritance and Terminology 21 Key Concepts Genetics is a science and specific terms are used. Make sure that you know and understand the following terms before you continue. Locus:
X-Sheet 4 Genetics: Inheritance and Terminology 21 Key Concepts Genetics is a science and specific terms are used. Make sure that you know and understand the following terms before you continue. Locus:
GENES AND CHROMOSOMES CHROMOSOMES IN SEX CELLS. Horse Science: How Inheritance Works in Horses Page 3. dam unite and grow into the new animal.
 Horse Science: How Inheritance Works in Horses Page 3 Two tiny cells are the only links of inheritance an animal has with its parents. A sperm cell from the sire and an egg cell from the dam unite and
Horse Science: How Inheritance Works in Horses Page 3 Two tiny cells are the only links of inheritance an animal has with its parents. A sperm cell from the sire and an egg cell from the dam unite and
Optimization of a crossing system using mate selection
 Genet. Sel. Evol. 38 (2006) 147-165 147 c INRA, EDP Sciences, 2006 DOI: 10.1051/gse:2005033 Original article Optimization of a crossing system using mate selection Yongjun Li a,b,juliush.j.vanderwerf a,brianp.kinghorn
Genet. Sel. Evol. 38 (2006) 147-165 147 c INRA, EDP Sciences, 2006 DOI: 10.1051/gse:2005033 Original article Optimization of a crossing system using mate selection Yongjun Li a,b,juliush.j.vanderwerf a,brianp.kinghorn
Use of Young and Proven Sires with Genomic Evaluations for Improving Milk Yield in Thai Multibreed Dairy Cattle
 Use of Young and Proven Sires with Genomic Evaluations for Improving Milk Yield in Thai Multibreed Dairy Cattle 1 Department of Animal Science, Faculty of Agriculture, Kasetsart University, 50 Ladyao,
Use of Young and Proven Sires with Genomic Evaluations for Improving Milk Yield in Thai Multibreed Dairy Cattle 1 Department of Animal Science, Faculty of Agriculture, Kasetsart University, 50 Ladyao,
Genetics and Punnett Squares
 Name: Class: Date: Genetics and Punnett Squares Introduction The results of crossing two individuals can be predicted by making a Punnett square. At the top of the square, the two possible alleles from
Name: Class: Date: Genetics and Punnett Squares Introduction The results of crossing two individuals can be predicted by making a Punnett square. At the top of the square, the two possible alleles from
Imputation of genotypes in Danish purebred and two-way crossbred pigs using low-density panels
 Xiang et al. Genetics Selection Evolution (2015) 47:54 DOI 10.1186/s12711-015-0134-4 Genetics Selection Evolution RESEARCH ARTICLE Open Access Imputation of genotypes in Danish purebred and two-way crossbred
Xiang et al. Genetics Selection Evolution (2015) 47:54 DOI 10.1186/s12711-015-0134-4 Genetics Selection Evolution RESEARCH ARTICLE Open Access Imputation of genotypes in Danish purebred and two-way crossbred
-- Results and Discussion
 Relationship of Beef Sire Birth Weight and Weaning Weight Expected Progeny Differences to Actual Performance of Crossbred Offspring SDSU M.B. Long1 and D.M. Marshall2 Department of Animal and Range Sciences
Relationship of Beef Sire Birth Weight and Weaning Weight Expected Progeny Differences to Actual Performance of Crossbred Offspring SDSU M.B. Long1 and D.M. Marshall2 Department of Animal and Range Sciences
Colour Genetics. Page 1 of 6. TinyBear Pomeranians CKC Registered Copyright All rights reserved.
 Colour Genetics Dogs have probably been domesticated for at least 15 000 years. The first dogs likely had an appearance much like the wolf or coyote. However, over the time they have been domesticated,
Colour Genetics Dogs have probably been domesticated for at least 15 000 years. The first dogs likely had an appearance much like the wolf or coyote. However, over the time they have been domesticated,
Selective Genotyping for Marker Assisted Selection Strategies for Soybean Yield Improvement. Ben Fallen
 Selective Genotyping for Marker Assisted Selection Strategies for Soybean Yield Improvement Ben Fallen INTRODUCTION! Plant biotechnology in plant breeding offers new possibilities for: increased productivity
Selective Genotyping for Marker Assisted Selection Strategies for Soybean Yield Improvement Ben Fallen INTRODUCTION! Plant biotechnology in plant breeding offers new possibilities for: increased productivity
ENHANCED PARKWAY STUDY: PHASE 2 CONTINUOUS FLOW INTERSECTIONS. Final Report
 Preparedby: ENHANCED PARKWAY STUDY: PHASE 2 CONTINUOUS FLOW INTERSECTIONS Final Report Prepared for Maricopa County Department of Transportation Prepared by TABLE OF CONTENTS Page EXECUTIVE SUMMARY ES-1
Preparedby: ENHANCED PARKWAY STUDY: PHASE 2 CONTINUOUS FLOW INTERSECTIONS Final Report Prepared for Maricopa County Department of Transportation Prepared by TABLE OF CONTENTS Page EXECUTIVE SUMMARY ES-1
Università degli Studi di Firenze
 The Design of In vivo Conservation Programmes In any population it is inevitable that some genetic variation will be lost over time Minimisation of the loss of genetic variation is equivalent to minimisation
The Design of In vivo Conservation Programmes In any population it is inevitable that some genetic variation will be lost over time Minimisation of the loss of genetic variation is equivalent to minimisation
Analysis of Estimated Breeding Values for Marble Score, Carcase Weight and the Terminal Carcase Index
 Analysis of Estimated Breeding Values for Marble Score, Carcase Weight and the Terminal Carcase Index Authors: 1. Matthew McDonagh CEO 2. Carel Teseling Technical Service Manager Australian Wagyu association
Analysis of Estimated Breeding Values for Marble Score, Carcase Weight and the Terminal Carcase Index Authors: 1. Matthew McDonagh CEO 2. Carel Teseling Technical Service Manager Australian Wagyu association
8. How many different kinds of gametes can normally be produced by an organism with the genotype RrYy? A) 1 B) 2 C) 3 D) 4
 Name: Biology 3201 Heredity Assignment March 25, 2011 Part A: Multiple choice. Choose the BEST answer. If you are unsure, guess. (15 points) 1. Which branch of biology deals with the principles of variation
Name: Biology 3201 Heredity Assignment March 25, 2011 Part A: Multiple choice. Choose the BEST answer. If you are unsure, guess. (15 points) 1. Which branch of biology deals with the principles of variation
Archival copy: for current recommendations see or your local extension office.
 NAME ADDRESS CLUB 4-H HORSE PROGRAM HORSE SCIENCE This educational material has been prepared for 4-H use by the Cooperative Extension Services of the U.S. Department of Agriculture and State Land-Grant
NAME ADDRESS CLUB 4-H HORSE PROGRAM HORSE SCIENCE This educational material has been prepared for 4-H use by the Cooperative Extension Services of the U.S. Department of Agriculture and State Land-Grant
Mission to Mars. Day 8 Heredity
 Mission to Mars Day 8 Heredity 7.14A define heredity as the passage of genetic instructions from one generation to the next generation 7.14C recognize that inherited traits of individuals are governed
Mission to Mars Day 8 Heredity 7.14A define heredity as the passage of genetic instructions from one generation to the next generation 7.14C recognize that inherited traits of individuals are governed
Genome mapping in salmonid fish
 Genome mapping in salmonid fish State of the art and perspectives Dr Karim Gharbi Kevlin/Smith Fellow in Aquatic Epidemiology and Immunogenomics Milestones First microsatellite map for rainbow trout First
Genome mapping in salmonid fish State of the art and perspectives Dr Karim Gharbi Kevlin/Smith Fellow in Aquatic Epidemiology and Immunogenomics Milestones First microsatellite map for rainbow trout First
Revealing the Past and Present of Bison Using Genome Analysis
 Revealing the Past and Present of Bison Using Genome Analysis David Forgacs, Rick Wallen, Lauren Dobson, Amy Boedeker, and James Derr July 5, 2017 1 Presentation outline 1. What can genetics teach us about
Revealing the Past and Present of Bison Using Genome Analysis David Forgacs, Rick Wallen, Lauren Dobson, Amy Boedeker, and James Derr July 5, 2017 1 Presentation outline 1. What can genetics teach us about
Fruit Fly Exercise 1- Level 1
 Name StarGenetics Fruit Fly Exercise 1- Level 1 Description of StarGenetics In this exercise you will use StarGenetics, a software tool that simulates mating experiments, to analyze the nature and mode
Name StarGenetics Fruit Fly Exercise 1- Level 1 Description of StarGenetics In this exercise you will use StarGenetics, a software tool that simulates mating experiments, to analyze the nature and mode
FIRE PROTECTION. In fact, hydraulic modeling allows for infinite what if scenarios including:
 By Phil Smith, Project Manager and Chen-Hsiang Su, PE, Senior Consultant, Lincolnshire, IL, JENSEN HUGHES A hydraulic model is a computer program configured to simulate flows for a hydraulic system. The
By Phil Smith, Project Manager and Chen-Hsiang Su, PE, Senior Consultant, Lincolnshire, IL, JENSEN HUGHES A hydraulic model is a computer program configured to simulate flows for a hydraulic system. The
Breeding poll cattle: the alternative to dehorning
 Breeding poll cattle: the alternative to dehorning John Henshall Group Leader, Genetics Genomics 10 May 2012 CSIRO LIVESTOCK INDUSTRIES Horns Undesirable bruising hide damage injury to the animal to other
Breeding poll cattle: the alternative to dehorning John Henshall Group Leader, Genetics Genomics 10 May 2012 CSIRO LIVESTOCK INDUSTRIES Horns Undesirable bruising hide damage injury to the animal to other
20 th Int. Symp. Animal Science Days, Kranjska gora, Slovenia, Sept. 19 th 21 st, 2012.
 20 th Int. Symp. Animal Science Days, Kranjska gora, Slovenia, Sept. 19 th 21 st, 2012. COBISS: 1.08 Agris category code: L10 Estimating Age of Admixture in a Cattle Population based on SNP Chip Data Anamarija
20 th Int. Symp. Animal Science Days, Kranjska gora, Slovenia, Sept. 19 th 21 st, 2012. COBISS: 1.08 Agris category code: L10 Estimating Age of Admixture in a Cattle Population based on SNP Chip Data Anamarija
Pedigree Analysis of Holstein Dairy Cattle Populations
 Pedigree Analysis of Holstein Dairy Cattle Populations C.Maltecca, F.Canavesi, G.Gandini, A.Bagnato. Department VSA, University of Milan, Italy ANAFI, Cremona, Italy Summary A pedigree analysis on the
Pedigree Analysis of Holstein Dairy Cattle Populations C.Maltecca, F.Canavesi, G.Gandini, A.Bagnato. Department VSA, University of Milan, Italy ANAFI, Cremona, Italy Summary A pedigree analysis on the
Queue analysis for the toll station of the Öresund fixed link. Pontus Matstoms *
 Queue analysis for the toll station of the Öresund fixed link Pontus Matstoms * Abstract A new simulation model for queue and capacity analysis of a toll station is presented. The model and its software
Queue analysis for the toll station of the Öresund fixed link Pontus Matstoms * Abstract A new simulation model for queue and capacity analysis of a toll station is presented. The model and its software
LAB. NATURAL SELECTION OF STRAWFISH
 Period Date LAB. NATURAL SELECTION OF STRAWFISH You have already been introduced to the idea that when an organism is selected by nature to survive, it then has the opportunity to reproduce and pass on
Period Date LAB. NATURAL SELECTION OF STRAWFISH You have already been introduced to the idea that when an organism is selected by nature to survive, it then has the opportunity to reproduce and pass on
Chapter 3 Mendelian Inheritance
 Chapter 3 Mendelian Inheritance I. Genes, Chromosomes, and Genotypes II. Germ Cells and Their Formation III. Formation of the Embryo IV.The Randomness of Inheritance V. Dominance and Epistasis VI.Sex-Related
Chapter 3 Mendelian Inheritance I. Genes, Chromosomes, and Genotypes II. Germ Cells and Their Formation III. Formation of the Embryo IV.The Randomness of Inheritance V. Dominance and Epistasis VI.Sex-Related
EBVS are LESSONS FROM THE ASBP / 2. EBVs of Bulls Entered in the ASBP have provided a reliable Prediction of the Performance of their Progeny
 EBVS are EBVs of Bulls Entered in the ASBP have provided a reliable Prediction of the Performance of their Progeny The Angus Sire Benchmarking Program (ASBP) has demonstrated that there is great potential
EBVS are EBVs of Bulls Entered in the ASBP have provided a reliable Prediction of the Performance of their Progeny The Angus Sire Benchmarking Program (ASBP) has demonstrated that there is great potential
Applications of Linear Algebra: Genetics Armeen Moshrefi, Sarah Rogers, Meghan Wolf, Jesus Zambrano Math 247: Jen-Mei Chang
 Applications of Linear Algebra: Genetics Armeen Moshrefi, Sarah Rogers, Meghan Wolf, Jesus Zambrano Math 247: Jen-Mei Chang Introduction Genetics is a branch of biology that deals with the mechanisms of
Applications of Linear Algebra: Genetics Armeen Moshrefi, Sarah Rogers, Meghan Wolf, Jesus Zambrano Math 247: Jen-Mei Chang Introduction Genetics is a branch of biology that deals with the mechanisms of
Impact of the Friesian POLLED Mutation on Milk Production Traits in Holstein Friesian
 Faculty of Agricultural and Nutritional Sciences Institute of Animal Breeding and Husbandry Impact of the Friesian POLLED Mutation on Milk Production Traits in Holstein Friesian EAAP 2016 67th Annual Meeting
Faculty of Agricultural and Nutritional Sciences Institute of Animal Breeding and Husbandry Impact of the Friesian POLLED Mutation on Milk Production Traits in Holstein Friesian EAAP 2016 67th Annual Meeting
Chapter 2: Traits and How They Change
 Chapter 2: Traits and How They Change Section 2: Genetics Heredity x Genetics Mendel s experiments Punnett Square For the test: study power point and textbook Important Info to remember : We inherit 2
Chapter 2: Traits and How They Change Section 2: Genetics Heredity x Genetics Mendel s experiments Punnett Square For the test: study power point and textbook Important Info to remember : We inherit 2
Maximizing genetic progress in the new age of genomics
 Veerkamp et al. Maximizing genetic progress in the new age of genomics R.H. Fourdraine AgSource Cooperative Services, 135 Enterprise Drive, Verona, WI 53593, United States Genetics have historically played
Veerkamp et al. Maximizing genetic progress in the new age of genomics R.H. Fourdraine AgSource Cooperative Services, 135 Enterprise Drive, Verona, WI 53593, United States Genetics have historically played
CHAPTER III RESULTS. sampled from 22 streams, representing 4 major river drainages in New Jersey, and 1 trout
 CHAPTER III RESULTS Genetic Diversity Genotypes at 13 microsatellite DNA loci were determined for 238 brook trout sampled from 22 streams, representing 4 major river drainages in New Jersey, and 1 trout
CHAPTER III RESULTS Genetic Diversity Genotypes at 13 microsatellite DNA loci were determined for 238 brook trout sampled from 22 streams, representing 4 major river drainages in New Jersey, and 1 trout
The Science of Maryland Agriculture
 Edition 3 (2016) GOAL STATEMENT: Students will learn how to predict plant and animal offspring traits or characteristics using genetics. Note: Students will need to have previous knowledge of basic genetics,
Edition 3 (2016) GOAL STATEMENT: Students will learn how to predict plant and animal offspring traits or characteristics using genetics. Note: Students will need to have previous knowledge of basic genetics,
Online Companion to Using Simulation to Help Manage the Pace of Play in Golf
 Online Companion to Using Simulation to Help Manage the Pace of Play in Golf MoonSoo Choi Industrial Engineering and Operations Research, Columbia University, New York, NY, USA {moonsoo.choi@columbia.edu}
Online Companion to Using Simulation to Help Manage the Pace of Play in Golf MoonSoo Choi Industrial Engineering and Operations Research, Columbia University, New York, NY, USA {moonsoo.choi@columbia.edu}
Occasionally white fleeced Ryeland sheep will produce coloured fleeced lambs.
 Genetic Testing Service available for Members Occasionally white fleeced Ryeland sheep will produce coloured fleeced lambs. The Society offers members a genetic testing service, carried out by Cardiff
Genetic Testing Service available for Members Occasionally white fleeced Ryeland sheep will produce coloured fleeced lambs. The Society offers members a genetic testing service, carried out by Cardiff
Proceedings of the World Congress on Genetics Applied to Livestock Production,
 Gain obtained by marker assisted selection in salmonids S. Kjøglum 1, N. Santi 1, J. Ødegård 1, S.A. Korsvoll 1, J.S. Torgersen 1, D. Cichero 2, M. Medina 2, H. Roa 3 & T. Moen 1 1 AquaGen AS, P.P. Box
Gain obtained by marker assisted selection in salmonids S. Kjøglum 1, N. Santi 1, J. Ødegård 1, S.A. Korsvoll 1, J.S. Torgersen 1, D. Cichero 2, M. Medina 2, H. Roa 3 & T. Moen 1 1 AquaGen AS, P.P. Box
Basic Mendelian Genetics & Color Genetics Basic Definitions Mendel demonstrated with corn that genes could be predictably combined.
 Basic Mendelian Genetics & Color Genetics Basic Definitions Mendel demonstrated with corn that genes could be predictably combined. For horses, there are 32 pairs of chromosomes which hold 2.7 billion
Basic Mendelian Genetics & Color Genetics Basic Definitions Mendel demonstrated with corn that genes could be predictably combined. For horses, there are 32 pairs of chromosomes which hold 2.7 billion
Operating Conditions Aligned Ship Design and Evaluation
 First International Symposium on Marine Propulsors smp 09, Trondheim, Norway, June 2009 Operating Conditions Aligned Ship Design and Evaluation Lars Greitsch, Georg Eljardt, Stefan Krueger Hamburg University
First International Symposium on Marine Propulsors smp 09, Trondheim, Norway, June 2009 Operating Conditions Aligned Ship Design and Evaluation Lars Greitsch, Georg Eljardt, Stefan Krueger Hamburg University
Population Analysis & Breeding and Transfer Plan
 Population Analysis & Breeding and Transfer Plan Cape Vulture (Gyps coprotheres) AZA Species Survival Plan Red Program AZA Species Survival Plan Coordinator Mike Maxcy, Los Angeles Zoo mike.maxcy@lacity.org
Population Analysis & Breeding and Transfer Plan Cape Vulture (Gyps coprotheres) AZA Species Survival Plan Red Program AZA Species Survival Plan Coordinator Mike Maxcy, Los Angeles Zoo mike.maxcy@lacity.org
Inheritance of Leaf Shape and Main Vein Color in Caladium 1
 ENH1006 Inheritance of Leaf Shape and Main Vein Color in Caladium 1 Z. Deng and B.K. Harbaugh 2 The ornamental value of caladiums (Caladium hortulanum Birdsey) depends, to a great extent, on leaf characteristics
ENH1006 Inheritance of Leaf Shape and Main Vein Color in Caladium 1 Z. Deng and B.K. Harbaugh 2 The ornamental value of caladiums (Caladium hortulanum Birdsey) depends, to a great extent, on leaf characteristics
Example: sequence data set wit two loci [simula
 Example: sequence data set wit two loci [simulated data] -- 1 Example: sequence data set wit two loci [simula MIGRATION RATE AND POPULATION SIZE ESTIMATION using the coalescent and maximum likelihood or
Example: sequence data set wit two loci [simulated data] -- 1 Example: sequence data set wit two loci [simula MIGRATION RATE AND POPULATION SIZE ESTIMATION using the coalescent and maximum likelihood or
A Comparison of Mating Systems Utilizing Duroc, Hampshire and Yorkshire Breeds for Swine Production. E. R. Wilson and R. K. Johnson.
 Swine BREEDING A Comparison of Mating Systems Utilizing Duroc, Hampshire and Yorkshire Breeds for Swine Production E. R. Wilson and R. K. Johnson Story in Brief This study utilized eight years of crossbreeding
Swine BREEDING A Comparison of Mating Systems Utilizing Duroc, Hampshire and Yorkshire Breeds for Swine Production E. R. Wilson and R. K. Johnson Story in Brief This study utilized eight years of crossbreeding
Characterization of two microsatellite PCR multiplexes for high throughput. genotyping of the Caribbean spiny lobster, Panulirus argus
 Characterization of two microsatellite PCR multiplexes for high throughput genotyping of the Caribbean spiny lobster, Panulirus argus Nathan K. Truelove 1, Richard F. Preziosi 1, Donald Behringer Jr 2,
Characterization of two microsatellite PCR multiplexes for high throughput genotyping of the Caribbean spiny lobster, Panulirus argus Nathan K. Truelove 1, Richard F. Preziosi 1, Donald Behringer Jr 2,
GENETICS 20 FEBRUARY 2013
 GENETICS 20 FEBRUARY 2013 Lesson Description In this lesson, we: Look at the definition of genetics. Discuss Mendel s concept of dominance & the law of segregation. Cover terminology of the genetic concepts.
GENETICS 20 FEBRUARY 2013 Lesson Description In this lesson, we: Look at the definition of genetics. Discuss Mendel s concept of dominance & the law of segregation. Cover terminology of the genetic concepts.
Chapter 2: Traits and How They Change
 Chapter 2: Traits and How They Change Section 2: Genetics Heredity x Genetics Mendel s experiments Punnett Square For the test: study the powerpoint and textbook 1)What is Heredity? 2)What is genetics?
Chapter 2: Traits and How They Change Section 2: Genetics Heredity x Genetics Mendel s experiments Punnett Square For the test: study the powerpoint and textbook 1)What is Heredity? 2)What is genetics?
6 Jon E. Hess 1, Nathan R. Campbell 1, Margaret F. Docker 2,
 1 Use of genotyping-by-sequencing data to develop a highthroughput and multi-functional set of genetic markers for conservation applications in Pacific lamprey 2 3 4 5 6 Jon E. Hess 1, Nathan R. Campbell
1 Use of genotyping-by-sequencing data to develop a highthroughput and multi-functional set of genetic markers for conservation applications in Pacific lamprey 2 3 4 5 6 Jon E. Hess 1, Nathan R. Campbell
The Science of Maryland Agriculture
 GOAL STATEMENT: Students will learn how to predict plant and animal offspring traits or characteristics using genetics. OBJECTIVES: Students will apply the concepts of genotype and phenotype to scenarios
GOAL STATEMENT: Students will learn how to predict plant and animal offspring traits or characteristics using genetics. OBJECTIVES: Students will apply the concepts of genotype and phenotype to scenarios
Working with Marker Maps Tutorial
 Working with Marker Maps Tutorial Release 8.2.0 Golden Helix, Inc. September 25, 2014 Contents 1. Overview 2 2. Create Marker Map from Spreadsheet 4 3. Apply Marker Map to Spreadsheet 7 4. Add Fields
Working with Marker Maps Tutorial Release 8.2.0 Golden Helix, Inc. September 25, 2014 Contents 1. Overview 2 2. Create Marker Map from Spreadsheet 4 3. Apply Marker Map to Spreadsheet 7 4. Add Fields
Full Name: Period: Heredity EOC Review
 Full Name: Period: 1 4 5 6 7 Heredity EOC Review Directions: For each genotype below, indicate whether it is a heterozygous (write: He) OR homozygous (write: Ho). 1. Tt BB DD ff tt dd dd Ff TT Bb bb FF
Full Name: Period: 1 4 5 6 7 Heredity EOC Review Directions: For each genotype below, indicate whether it is a heterozygous (write: He) OR homozygous (write: Ho). 1. Tt BB DD ff tt dd dd Ff TT Bb bb FF
Population analysis and conservation options of the Norwegian Lundehund
 Population analysis and conservation options of the Norwegian Lundehund A. Kettunen and P. Berg NordGen, Nordic Genetic Resource Center M. Daverdin NTNU Museum, Norwegian University of Science and Technology
Population analysis and conservation options of the Norwegian Lundehund A. Kettunen and P. Berg NordGen, Nordic Genetic Resource Center M. Daverdin NTNU Museum, Norwegian University of Science and Technology
 57th Annual EAAP Meeting in Antalya, Turkey Session 6: Impact of reproduction technology on horse breeding programmes H6.2 Cloning and embryo transfer in selection plans in horses Ricard, A. and C. Dubois
57th Annual EAAP Meeting in Antalya, Turkey Session 6: Impact of reproduction technology on horse breeding programmes H6.2 Cloning and embryo transfer in selection plans in horses Ricard, A. and C. Dubois
Mendel s Second Set of Experiments Dihybrid Crosses
 Mendel s Second Set of Experiments Dihybrid Crosses Mendel s first set of experiments proved the Law of Segregation; an individual contains factors which segregate from one another during meiosis. Each
Mendel s Second Set of Experiments Dihybrid Crosses Mendel s first set of experiments proved the Law of Segregation; an individual contains factors which segregate from one another during meiosis. Each
59 th EAAP Annual Meeting August 24-27, 2008, Vilnius, Lithuania Abstract # 2908, Session 08
 59 th EAAP Annual Meeting August 24-27, 2008, Vilnius, Lithuania Abstract # 2908, Session 08 Genetic variability of the Skyros pony breed and its relationship with other Greek and foreign horse breeds
59 th EAAP Annual Meeting August 24-27, 2008, Vilnius, Lithuania Abstract # 2908, Session 08 Genetic variability of the Skyros pony breed and its relationship with other Greek and foreign horse breeds
Teams *List of Student Teams will be placed here * 4 Teams
 What is Science? Game Rules 1. Group member to answer question will rotate for each group response. 2. Groups will have a chance to steal the question once the time has gone off. 3. Use your buzzers to
What is Science? Game Rules 1. Group member to answer question will rotate for each group response. 2. Groups will have a chance to steal the question once the time has gone off. 3. Use your buzzers to
Unit 8: GENETICS PACKET
 Unit 8: GENETICS PACKET This packet is designed to help you understand several concepts about GENETICS. As you practice the exercises on each handout, you will be able to: Make and defend a claim based
Unit 8: GENETICS PACKET This packet is designed to help you understand several concepts about GENETICS. As you practice the exercises on each handout, you will be able to: Make and defend a claim based
Aim of breeding work
 Breeding in Denmark registrations, indices and breeding scheme International Limousin Congress 2012, Technical Meeting Hotel Radisson, Denmark Wednesday 4 th July 2012 Anders Fogh Aim of breeding work
Breeding in Denmark registrations, indices and breeding scheme International Limousin Congress 2012, Technical Meeting Hotel Radisson, Denmark Wednesday 4 th July 2012 Anders Fogh Aim of breeding work
Biology- Genetics: Who Dares Wins Probability and Heredity Lab Report
 iology- Genetics: Who Dares Wins Proaility and Heredity La Report QUESTION/PROLEM: How can you predict the possile results of genetic crosses? ACKGROUND INFO The purpose of this La report is to find out
iology- Genetics: Who Dares Wins Proaility and Heredity La Report QUESTION/PROLEM: How can you predict the possile results of genetic crosses? ACKGROUND INFO The purpose of this La report is to find out
Single-step genomic BLUP for national beef cattle evaluation in US:
 Single-step genomic BLUP for national beef cattle evaluation in US: from initial developments to final implementation Daniela Lourenco S. Tsuruta, B.O. Fragomeni, Y. Masuda, I. Aguilar A. Legarra, S. Miller,
Single-step genomic BLUP for national beef cattle evaluation in US: from initial developments to final implementation Daniela Lourenco S. Tsuruta, B.O. Fragomeni, Y. Masuda, I. Aguilar A. Legarra, S. Miller,
1) A child is born with blue eyes even though BOTH his parents have brown eyes. How is this possible?
 Warm Up 1) A child is born with blue eyes even though BOTH his parents have brown eyes. How is this possible? 2) Can two blue eyed parents have a brown eyed child? Explain why or why not. Genetics: The
Warm Up 1) A child is born with blue eyes even though BOTH his parents have brown eyes. How is this possible? 2) Can two blue eyed parents have a brown eyed child? Explain why or why not. Genetics: The
Genetics and Miniature horses. Munro Marx Unistel Medical laboratories
 Genetics and Miniature horses Munro Marx Unistel Medical laboratories Miniature Horses Miniature horses are bred all over the world They are particularly popular in Europe and the USA Over the last number
Genetics and Miniature horses Munro Marx Unistel Medical laboratories Miniature Horses Miniature horses are bred all over the world They are particularly popular in Europe and the USA Over the last number
57th Annual EAAP Meeting, Antalya, September
 G3.10 corresponding author: pariset@unitus.it @unitus.it. Genetic characterization by single nucleotide polymorphisms (SNPs) of Hungarian Grey and Maremmana cattle breeds L. Pariset 1, C. Marchitelli 1,
G3.10 corresponding author: pariset@unitus.it @unitus.it. Genetic characterization by single nucleotide polymorphisms (SNPs) of Hungarian Grey and Maremmana cattle breeds L. Pariset 1, C. Marchitelli 1,
Genetic analysis of radio-tagged westslope cutthroat trout from St. Mary s River and Elk River. April 9, 2002
 Genetic analysis of radio-tagged westslope cutthroat trout from St. Mary s River and Elk River April 9, 2002 Report prepared for: Angela Prince, M.Sc., R.P. Bio Westslope Fisheries 517 13 th Avenue South
Genetic analysis of radio-tagged westslope cutthroat trout from St. Mary s River and Elk River April 9, 2002 Report prepared for: Angela Prince, M.Sc., R.P. Bio Westslope Fisheries 517 13 th Avenue South
CAM Final Report John Scheele Advisor: Paul Ohmann I. Introduction
 CAM Final Report John Scheele Advisor: Paul Ohmann I. Introduction Herds are a classic complex system found in nature. From interactions amongst individual animals, group behavior emerges. Historically
CAM Final Report John Scheele Advisor: Paul Ohmann I. Introduction Herds are a classic complex system found in nature. From interactions amongst individual animals, group behavior emerges. Historically
MEMORANDUM. To: PRL Performance Standards Subgroup From: Donna Pratt Subject: Performance Method Recommendation Date: January 18, 2001
 MEMORANDUM To: PRL Performance Standards Subgroup From: Donna Pratt Subject: Performance Method Recommendation Date: January 18, 2001 The Emergency Demand Response Program Manual is in its final phase
MEMORANDUM To: PRL Performance Standards Subgroup From: Donna Pratt Subject: Performance Method Recommendation Date: January 18, 2001 The Emergency Demand Response Program Manual is in its final phase
Genetics Test Review
 Genetics Test Review Vocabulary to Know 1) Punnett Square chart used to determine the probable outcome of genetic crosses 2) Heredity The passing of traits from parent to offspring 3) Gregor Mendel The
Genetics Test Review Vocabulary to Know 1) Punnett Square chart used to determine the probable outcome of genetic crosses 2) Heredity The passing of traits from parent to offspring 3) Gregor Mendel The
SWINE BREEDING SYSTEMS
 Extension Bulletin E-3107 New August 2010 SWINE BREEDING SYSTEMS for Alternative Pork Chains: Breeding Programs Ronald O. Bates State Swine Specialist Michigan State University Introduction The pork industry
Extension Bulletin E-3107 New August 2010 SWINE BREEDING SYSTEMS for Alternative Pork Chains: Breeding Programs Ronald O. Bates State Swine Specialist Michigan State University Introduction The pork industry
Genetic Trend for Milk. U.S. dairy population and milk yield
 52nd Florida Dairy Production Conference Opportunity Costs When Choosing Service Sires 2016 Florida Dairy Production Conference University of Florida April 6, 2016 Dr. Ole Meland Chairman of the Council
52nd Florida Dairy Production Conference Opportunity Costs When Choosing Service Sires 2016 Florida Dairy Production Conference University of Florida April 6, 2016 Dr. Ole Meland Chairman of the Council
Gregor Mendel. 19 th century Austrian monk Studied pea plants in his garden. Father of modern genetics
 GENETICS Gregor Mendel 19 th century Austrian monk Studied pea plants in his garden Father of modern genetics Blending Theory Incorrect idea that was popular during Mendel s time Thought that an offspring
GENETICS Gregor Mendel 19 th century Austrian monk Studied pea plants in his garden Father of modern genetics Blending Theory Incorrect idea that was popular during Mendel s time Thought that an offspring
NON-MENDELIAN GENETICS:
 NON-MENDELIAN GENETICS: INCOMPLETE DOMIANCE CO-DOMINANCE (aka Multiple Alleles) SEX-LINKED INHERITANCE POLYGENIC TRAITS Name: This practice contains assignments for the standards listed above. We will
NON-MENDELIAN GENETICS: INCOMPLETE DOMIANCE CO-DOMINANCE (aka Multiple Alleles) SEX-LINKED INHERITANCE POLYGENIC TRAITS Name: This practice contains assignments for the standards listed above. We will
Test Results Summary. Owner: Darby Dan Farm 8 th Nov Distance Plus (USA) v1.0. Distance Plus (SAF) v1.0. Distance Plus (ANZ) v1.
 Test Results Summary Owner: Darby Dan Farm 8 th Nov 2017 Sample ID Horse Name Sire Dam Sex Year of Birth Speed Gene Test (ANZ) (EU) (USA) (SAF) Dirt V Turf Elite Performance Test v3.1 Inbreeding Value
Test Results Summary Owner: Darby Dan Farm 8 th Nov 2017 Sample ID Horse Name Sire Dam Sex Year of Birth Speed Gene Test (ANZ) (EU) (USA) (SAF) Dirt V Turf Elite Performance Test v3.1 Inbreeding Value
September Population analysis of the Finnish Spitz breed
 Population analysis of the Finnish Spitz breed Genetic analysis of the Kennel Club pedigree records of the UK Finnish Spitz population has been carried out with the aim of estimating the rate of loss of
Population analysis of the Finnish Spitz breed Genetic analysis of the Kennel Club pedigree records of the UK Finnish Spitz population has been carried out with the aim of estimating the rate of loss of
Introduction to Genetics
 Name: Introduction to Genetics Keystone Assessment Anchor: BIO.B.2.1.1: Describe and/or predict observed patterns of inheritance (i.e. dominant, recessive, co-dominance, incomplete dominance, sex-linked,
Name: Introduction to Genetics Keystone Assessment Anchor: BIO.B.2.1.1: Describe and/or predict observed patterns of inheritance (i.e. dominant, recessive, co-dominance, incomplete dominance, sex-linked,
CALF PERFORMANCE AND COW WEIGHT AND CONDITION FOR COWS SIRED BY HIGH AND LOW MILK EPD ANGUS AND POLLED HEREFORD BULLS
 CALF PERFORMANCE AND COW WEIGHT AND CONDITION FOR COWS SIRED BY HIGH AND LOW MILK EPD ANGUS AND POLLED HEREFORD BULLS D.S. Buchanan 1, R. Gosz 2, E. Nesamvuni 2 and L. Knori 3 Story in Brief Calf birth
CALF PERFORMANCE AND COW WEIGHT AND CONDITION FOR COWS SIRED BY HIGH AND LOW MILK EPD ANGUS AND POLLED HEREFORD BULLS D.S. Buchanan 1, R. Gosz 2, E. Nesamvuni 2 and L. Knori 3 Story in Brief Calf birth
September Population analysis of the Portuguese Water Dog breed
 Population analysis of the Portuguese Water Dog breed Genetic analysis of the Kennel Club pedigree records of the UK Portuguese Water Dog population has been carried out with the aim of estimating the
Population analysis of the Portuguese Water Dog breed Genetic analysis of the Kennel Club pedigree records of the UK Portuguese Water Dog population has been carried out with the aim of estimating the
Torpedoes on Target: EAs on Track
 113 Torpedoes on Target: EAs on Track Nicole Patterson, Luigi Barone and Cara MacNish School of Computer Science & Software Engineering The University of Western Australia M002, 35 Stirling Highway, Crawley,
113 Torpedoes on Target: EAs on Track Nicole Patterson, Luigi Barone and Cara MacNish School of Computer Science & Software Engineering The University of Western Australia M002, 35 Stirling Highway, Crawley,
The Sustainability of Atlantic Salmon (Salmo salar L.) in South West England
 The Sustainability of Atlantic Salmon (Salmo salar L.) in South West England Submitted by Sarah-Louise Counter to the University of Exeter as a thesis for the degree of Doctor of Philosophy in Biological
The Sustainability of Atlantic Salmon (Salmo salar L.) in South West England Submitted by Sarah-Louise Counter to the University of Exeter as a thesis for the degree of Doctor of Philosophy in Biological
POWER Quantifying Correction Curve Uncertainty Through Empirical Methods
 Proceedings of the ASME 2014 Power Conference POWER2014 July 28-31, 2014, Baltimore, Maryland, USA POWER2014-32187 Quantifying Correction Curve Uncertainty Through Empirical Methods ABSTRACT Christopher
Proceedings of the ASME 2014 Power Conference POWER2014 July 28-31, 2014, Baltimore, Maryland, USA POWER2014-32187 Quantifying Correction Curve Uncertainty Through Empirical Methods ABSTRACT Christopher
Population structure and genetic diversity of Bovec sheep from Slovenia preliminary results
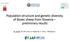 Population structure and genetic diversity of Bovec sheep from Slovenia preliminary results M. Simčič, M. Žan Lotrič, D. Bojkovski, E. Ciani, I. Medugorac Introduction Jezersko Solčava sheep Bovec sheep
Population structure and genetic diversity of Bovec sheep from Slovenia preliminary results M. Simčič, M. Žan Lotrič, D. Bojkovski, E. Ciani, I. Medugorac Introduction Jezersko Solčava sheep Bovec sheep
Legendre et al Appendices and Supplements, p. 1
 Legendre et al. 2010 Appendices and Supplements, p. 1 Appendices and Supplement to: Legendre, P., M. De Cáceres, and D. Borcard. 2010. Community surveys through space and time: testing the space-time interaction
Legendre et al. 2010 Appendices and Supplements, p. 1 Appendices and Supplement to: Legendre, P., M. De Cáceres, and D. Borcard. 2010. Community surveys through space and time: testing the space-time interaction
September Population analysis of the Dogue De Bordeaux breed
 Population analysis of the Dogue De Bordeaux breed Genetic analysis of the Kennel Club pedigree records of the UK Dogue De Bordeaux population has been carried out with the aim of estimating the rate of
Population analysis of the Dogue De Bordeaux breed Genetic analysis of the Kennel Club pedigree records of the UK Dogue De Bordeaux population has been carried out with the aim of estimating the rate of
Identification of the dwarfing gene Dw1
 Identification of the dwarfing gene Dw1 Toshi M. Foster, Jean-Marc Celton, David Chagné, and Susan E. Gardiner Dwarfing rootstocks reduce scion vigour Less branching More flowers Earlier shoot termination
Identification of the dwarfing gene Dw1 Toshi M. Foster, Jean-Marc Celton, David Chagné, and Susan E. Gardiner Dwarfing rootstocks reduce scion vigour Less branching More flowers Earlier shoot termination
At each type of conflict location, the risk is affected by certain parameters:
 TN001 April 2016 The separated cycleway options tool (SCOT) was developed to partially address some of the gaps identified in Stage 1 of the Cycling Network Guidance project relating to separated cycleways.
TN001 April 2016 The separated cycleway options tool (SCOT) was developed to partially address some of the gaps identified in Stage 1 of the Cycling Network Guidance project relating to separated cycleways.
System Operating Limit Definition and Exceedance Clarification
 System Operating Limit Definition and Exceedance Clarification The NERC defined term System Operating Limit (SOL) is used extensively in the NERC Reliability Standards; however, there is much confusion
System Operating Limit Definition and Exceedance Clarification The NERC defined term System Operating Limit (SOL) is used extensively in the NERC Reliability Standards; however, there is much confusion
Value of Black Hereford Registration
 Value of Black Hereford Registration Increase Marketability Whether one owns 10 or 500 Black Hereford cows, a registration certificate shows other producers that you are willing to take the time to document
Value of Black Hereford Registration Increase Marketability Whether one owns 10 or 500 Black Hereford cows, a registration certificate shows other producers that you are willing to take the time to document
Contemporary Grouping for Beef Cattle Genetic Evaluation
 Contemporary Grouping for Beef Cattle Genetic Evaluation Reprinted from the 8 th Edition of the Beef Improvement Federation Guidelines Every weight or measurement of an animal is an observation of its
Contemporary Grouping for Beef Cattle Genetic Evaluation Reprinted from the 8 th Edition of the Beef Improvement Federation Guidelines Every weight or measurement of an animal is an observation of its
